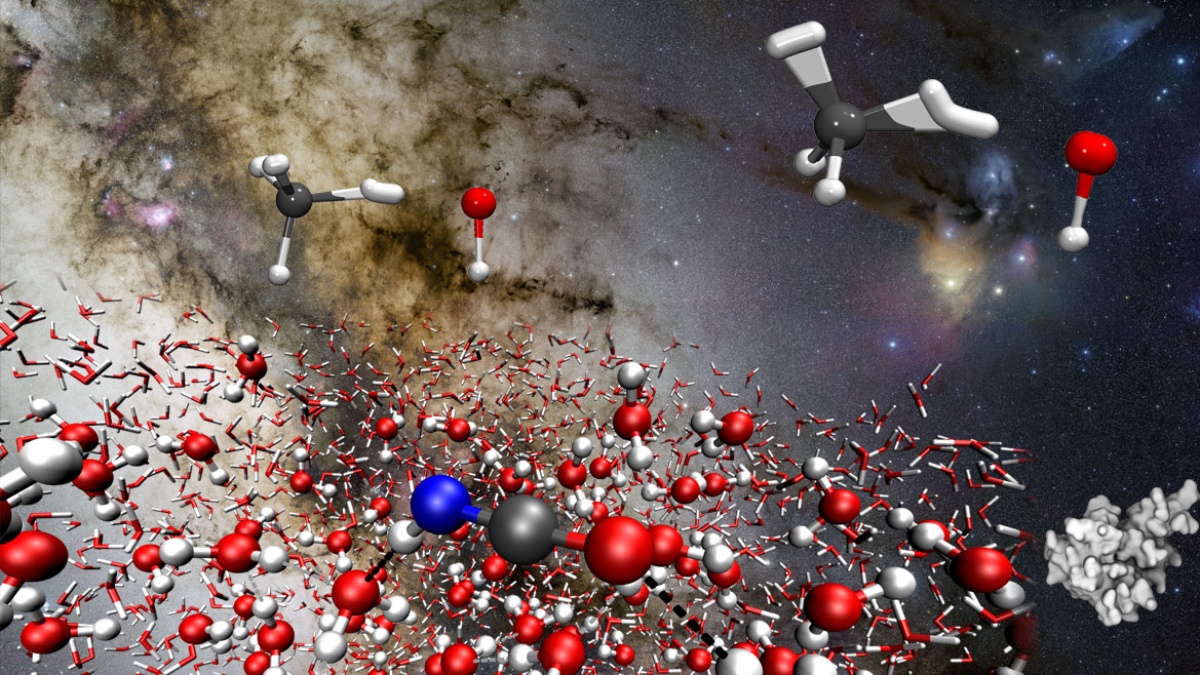
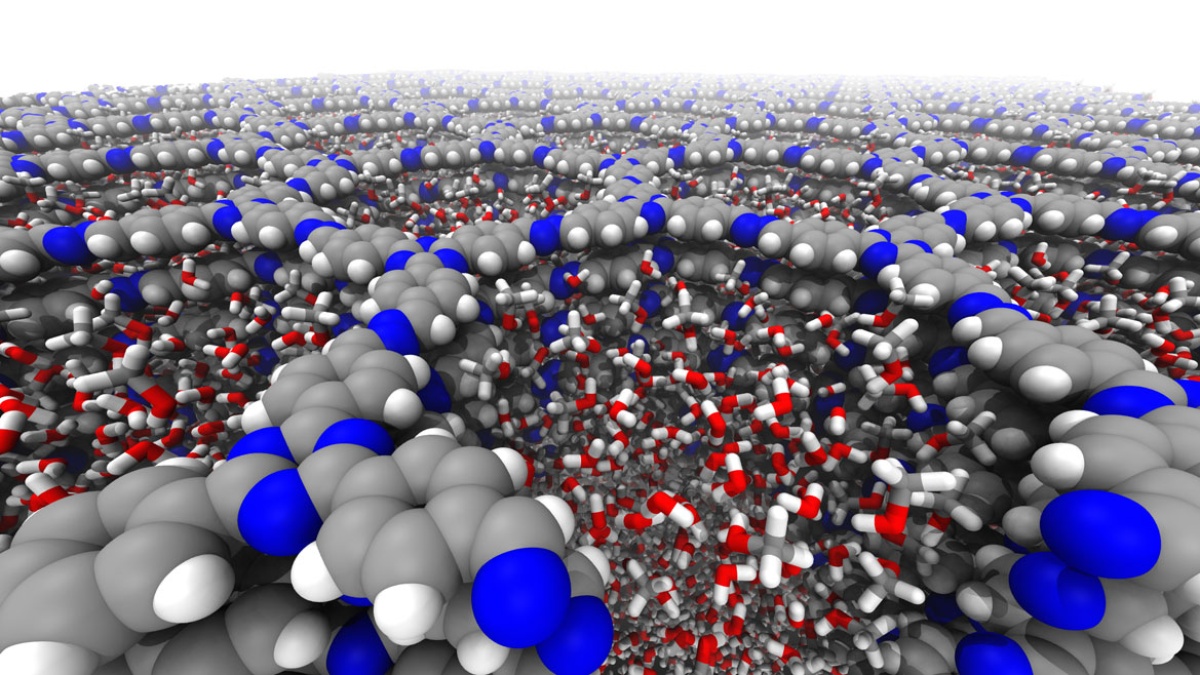
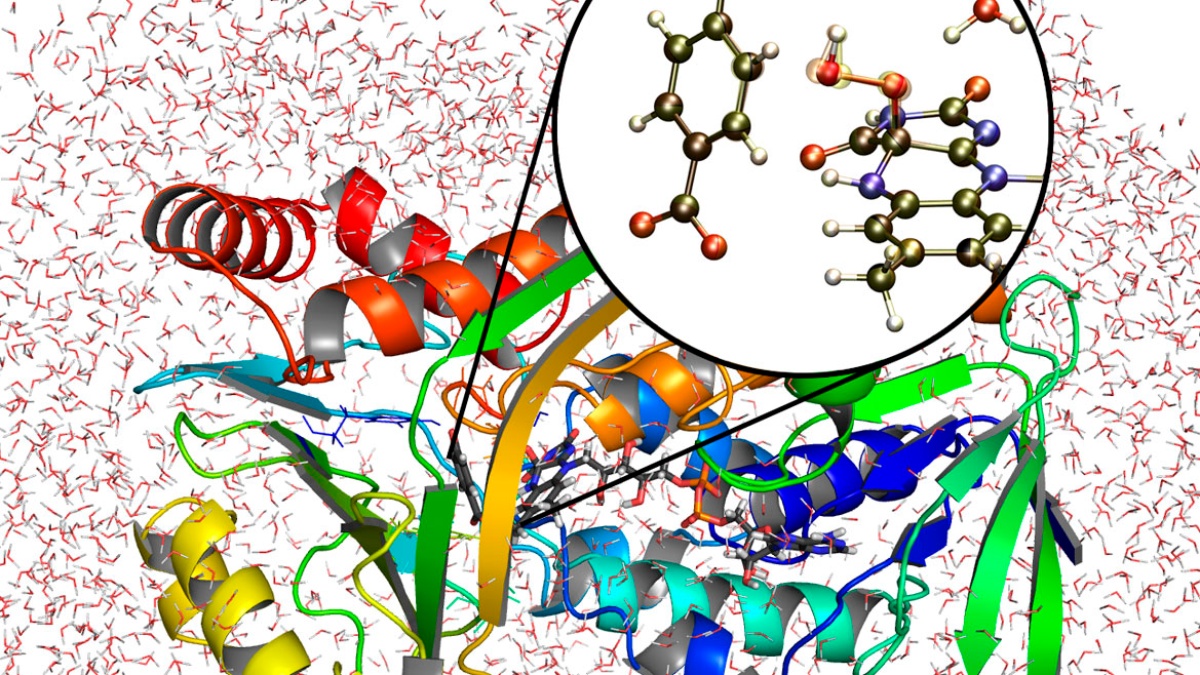
The Computational Chemistry Group simulates chemical and biochemical reactions under consideration of environmental effects. We complement experiments and provide answers where experiments alone cannot do so. We study catalysis, surface processes, astrochemistry, enzymes, and materials.
Application areas:
- Catalysis
- Astrochemistry
- Surface science (CRC ATLAS)
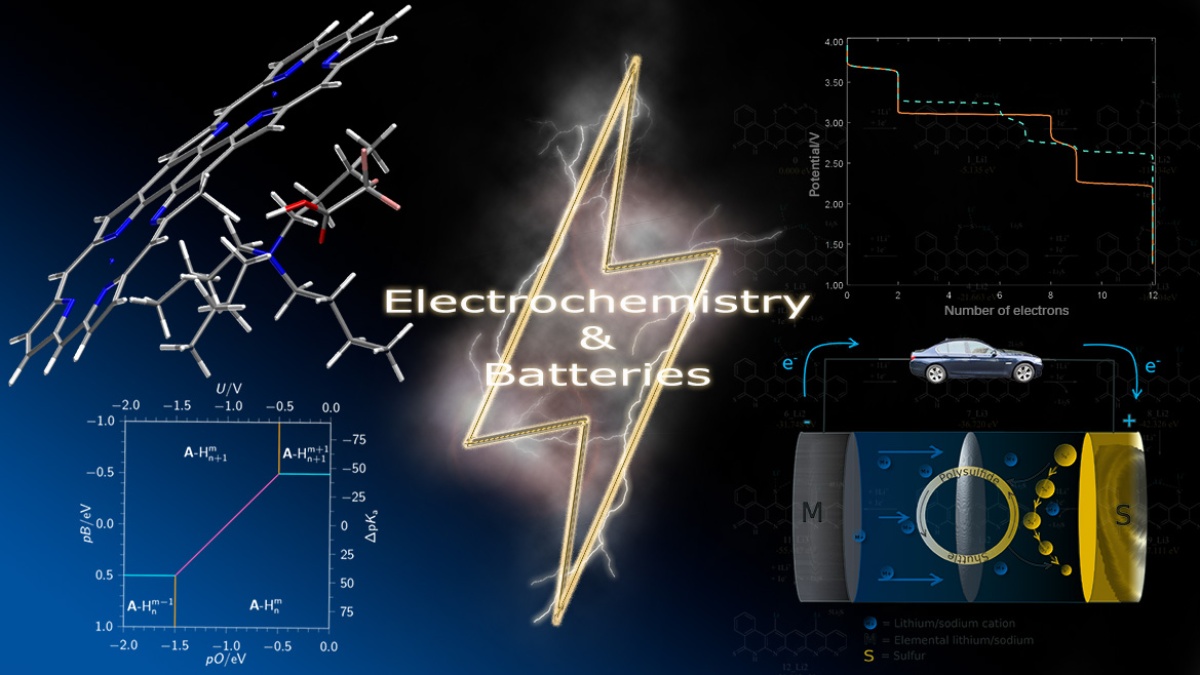
- Electrochemistry, batteries
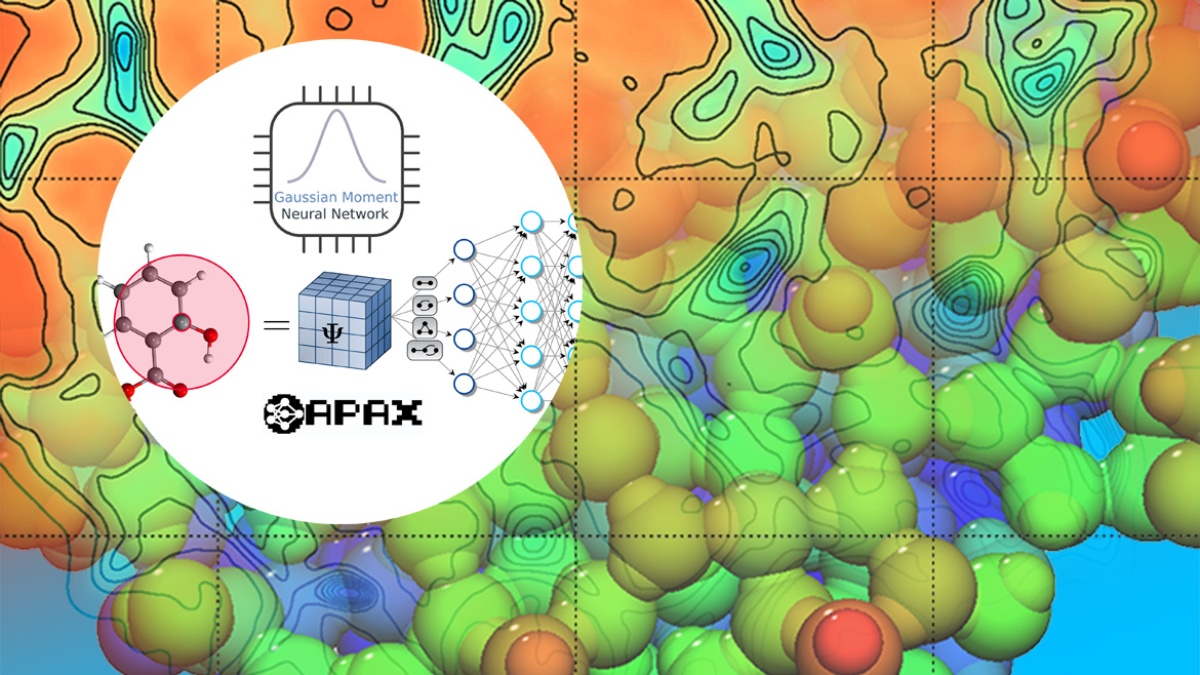
Method development:
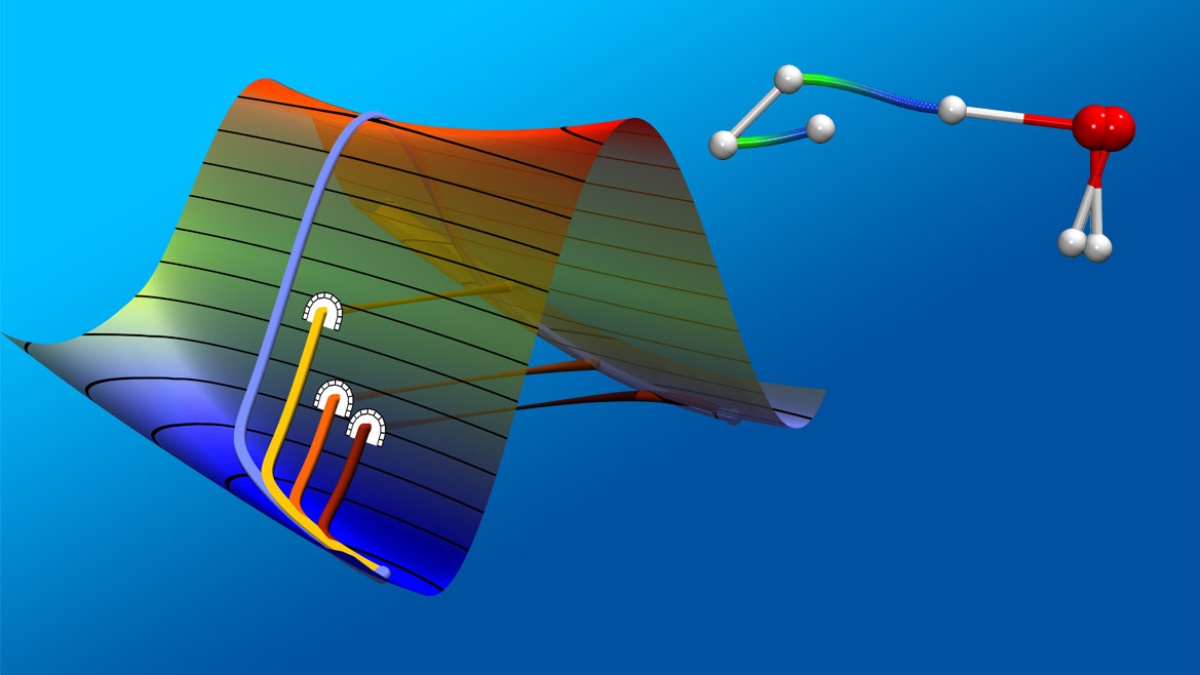
We develop methods that allow us to investigate these kinds of problems in innovative ways. We use machine learning techniques to investigate chemical reactions and to optimize geometries. We co-author the program package ChemShell, and we lead the development of the optimization library DL-FIND. Our group extends the capabilities of geometry optimization in systems with many thousand degrees of freedom and developed methods to calculate tunneling processes in large systems as well as the free-energy sampling technique umbrella integration.
Main Funding Sources
- ERC Consolidator Grant TUNNELCHEM
Completed grant by the European Research Council - SimTech Cluster of Excellence
A project that brought the PI to Stuttgart and that keeps us inspired! - SFB 1333: Molecular Heterogeneous Catalysis in Confined Geometries
Opening new frontiers in catalysis - LILAC consortium in AstroChemistry
Astrochemistry requires close interaction with astronomical observation, lab astrophysics, and modeling - Artificial Intelligence Software Academy
Cutting-edge research and training in AI - SPP 2363 on Utilization and Development of Machine Learning for Molecular Applications – Molecular Machine Learning
AI and ML for molecular applications - SFB 1667: Advancing Technologies of Very Low Altitude Satellites
We find out what the remaining atmosphere in 200 km altitude does to satellites.